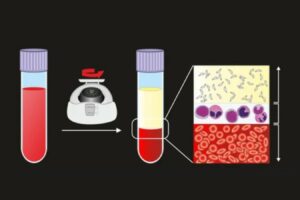
For researchers working in immunology and drug development, the human leukocyte antigen (HLA) system or complex is a critical element to keep in mind when planning experiments.
HLA information is used for a variety of purposes in preclinical research such as drug screening, determining disease susceptibility or inducing T cell stimulation in culture. At Cytologics, we frequently have customers ask questions such as:
- “Do I really need high resolution HLA typing on my research samples?”
- “Will low resolution typing provide enough accuracy and detail for my experiment?”
Although the HLA system is central to adaptive immunity and cell therapies, many scientists who do not specialize in the field find the nomenclature and typing techniques confusing.
Let’s take a look at the various HLA typing methods and determine which is the best fit for your research.
HLA Overview
One of the main uses of HLA typing is for identifying the best recipient and donor match prior to hematopoietic stem cell transplantation. However, researchers also use HLA data in a wide range of drug development applications.
As a regulator of the immune system, HLA can predict immune responses to various infectious diseases, autoimmune conditions and genetic disorders. [1] For instance, patients with HLA-B*15:01 have been associated with specific cancer responses to checkpoint blockade immunotherapy. [2]
The advantage of working with HLA-typed cells is that these samples provide an optimal in vitro control system to investigate immune-mediated disease and adverse drug reactions. Samples with HLA typing provide more reliable models than animal cells, therefore transferability to clinical use is more assured.
It’s important to note that HLA genes are the most polymorphic gene family found in the entire human genome with more than 30,000 different HLA alleles reported since the discovery of the HLA genes in the early 1950s. [3]
Due to the ever increasing number of alleles discovered, the methods and technologies to characterize HLA gene sequences with full accuracy is constantly evolving.
Typing technologies have progressed from simple serologic methods to molecular analysis and next-generation sequencing (NGS) which have improved patient outcomes following transplantation.
What Exactly Is the Difference between Low Resolution and High Resolution HLA Typing?
HLA typing methods are generally characterized at low, intermediate and high resolution depending on their power to discriminate between HLA alleles.
Low Resolution – an antigen-level result composing the first field of the HLA nomenclature. Examples include: A01; A02. If the resolution corresponds to a serologic equivalent, this typing result should also be called low resolution. [4]
Intermediate Resolution – a DNA-based typing result that includes a subset of alleles sharing the digits in the first field of their allele name and that excludes some alleles sharing this field. [5]
High Resolution – defined as a set of alleles that specify and encode the same protein sequence within the antigen recognition site. [6] High resolution typing can also consist of a set of alleles that specify and encode the same protein sequence for the antigen binding domain of an HLA molecule and that excludes alleles that are not expressed as cell-surface proteins. This level of resolution resolves all ambiguities resulting from polymorphisms located within exons 2 and 3 for class I loci, and within exon 2 for class II loci.
Which HLA Typing Method is Best for My Application?
Currently, the standard of care for HLA typing to select a donor for transplantation is to use a low or intermediate resolution method first, then type eligible candidates using a high-resolution method to confirm results and select the best matched donor.
HLA resolution is also a key consideration in preclinical research involving immune responses to disease. HLA molecules present foreign antigens to elicit T cell responses, so the number of distinct HLA allotypes expressed on the cell surface is directly related to the range of foreign antigens. [7]
The decision on whether to use a high, intermediate or low resolution HLA typing method for your research will be driven by the required turnaround time and cost. The table below compares various typing methods and resolutions:
Table 1. Comparison of HLA Typing Methods
| Method | Resolution | Advantages | Disadvantages |
|---|---|---|---|
| Serological | Low | – Fast turnaround – Low cost – Adjunct typing method |
– Requires live cells – Provides only antigen-level detail – Inadequate for HSC transplant matching |
| PCR with sequence specific primers (PCR-SSP) | Intermediate | – Very rapid test (3-4 hours) – DNA-based HLA typing |
– Limited resolution – Only amplifies specific HLA alleles |
| Sequence specific oligonucleotide probing (PCR-SSOP) | Intermediate | – Low cost – High throughput – Ability to determine multiple locus types |
– Time consuming – Limited resolution |
| Sanger sequencing-based typing (SBT) | High | – High accuracy – Simple data analysis – Established workflow – Ideal for low target numbers |
– Time-consuming protocols – Low throughput – Unphased data – Genotype ambiguity |
| Next-generation Sequencing (NGS) | High | – High accuracy – High sensitivity – High throughput – Reduced genotype ambiguity |
– High cost – Long sequencing time – Highly complex method |
Conclusion
Although the most significant impact of HLA typing is the selection of donors for transplantation, this technology is also a valuable tool for cell therapy development and immunology studies.
In fact, the implications of HLA are now being considered in different fields of research including oncology, cardiology, dermatology and infectious disease, among others.
Drug discovery experiments in these fields may require donor or patient samples with well-defined HLA types. The good news is that you can avoid time consuming and costly donor screenings to spot specific HLA types by working with commercial providers like Cytologics.
We have a comprehensive collection of HLA-typed human primary cells readily available from a diverse donor base. We offer customers the ability to select samples based on demographic and physical characteristics and HLA type so you can optimize your research.
For more information about our HLA-typed samples, please contact us or check out our related blog posts on the HLA system.
References
[1] Crux NB, Elahi S. Human Leukocyte Antigen (HLA) and Immune Regulation: How Do Classical and Non-Classical HLA Alleles Modulate Immune Response to Human Immunodeficiency Virus and Hepatitis C Virus Infections? Front Immunol. 2017 Jul 18;8:832. doi: 10.3389/fimmu.2017.00832. PMID: 28769934; PMCID: PMC5513977.
[2] Chowell D, et al. Patient HLA class I genotype influences cancer response to checkpoint blockade immunotherapy. Science 10.1126/science.aao4572 (2017).
[3] Robinson J, Halliwell JA, Hayhurst JH, Flicek P, Parham P, Marsh SGE. The IPD and IMGT/HLA database: allele variant databases. Nucleic Acids Research (2015) 43:D423-431 HLA Nomenclature @ hla.alleles.org.
[4] Sanchez-Mazas A, et al. Strategies to work with HLA data in human populations for histocompatibility, clinical transplantation, epidemiology and population genetics: HLA-NET methodological recommendations. Int. J. Immunogenetics (2012) 39(6) 459–476.
[5] Nunes E, et al. Definitions of histocompatibility typing terms. Blood (2011) 118 (23): e180–e183. https://doi.org/10.1182/blood-2011-05-353490.
[6] Nunes JM, Buhler S, Roessli D, Sanchez-Mazas A and the HLA-net 2013 collaboration (2014) The HLA-net GENE[RATE] pipeline for effective HLA data analysis and its application to 145 population samples from Europe and neighbouring areas. Tissue Antigens 83: 307-323.
[7] Juanes-Velasco P, et al. Deciphering Human Leukocyte Antigen Susceptibility Maps From Immunopeptidomics Characterization in Oncology and Infections. Front Cell Infect Microbiol. 2021 May 28;11:642583. doi: 10.3389/fcimb.2021.642583. PMID: 34123866; PMCID: PMC8195621.